Présentation technologique utilisant la spectrométrie de masse haute résolution (HRMS)
Notre approche technologique pour l'identification et la quantification des protéines peut être divisée en deux stratégies : une approche globale, basée sur la découverte, et une approche plus ciblée, fondée sur des hypothèses.
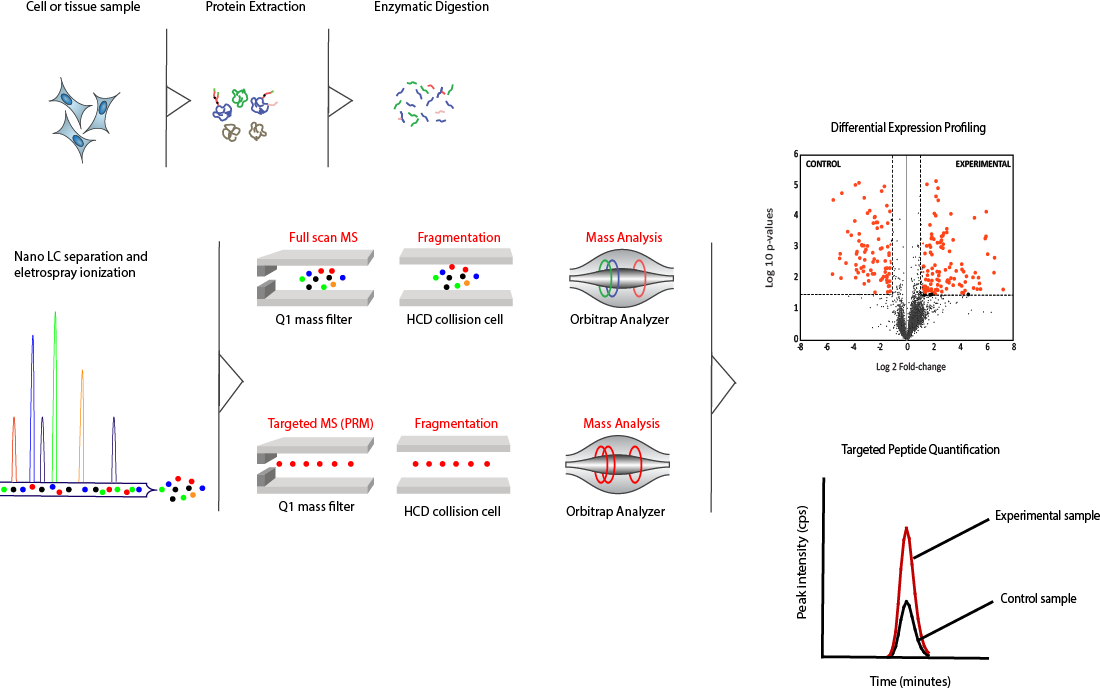
Les protéines des tissus ou des cultures cellulaires sont d'abord extraites puis digérées par voie enzymatique pour générer un mélange complexe de peptides. Ce mélange est d'abord séparé chromatographiquement à l'aide de colonnes capillaires nanométriques, puis transféré en phase gazeuse par ionisation par électrospray.
Dans la stratégie de détection basée sur la découverte, tous les ions peptidiques sont détectés et séquencés par l'analyseur de masse Orbitrap. Cela nous fournit un profil global de tous les composants présents dans notre mélange peptidique en utilisant la LC-MS à balayage complet. Pour se concentrer sur un sous-ensemble plus petit de protéines d'intérêt, une stratégie de détection alternative (surveillance des réactions parallèles, ou PRM) utilisant une liste cible de peptides est appliquée.
Les peptides peuvent ensuite être identifiés et quantifiés à partir des intensités de leurs spectres de masse correspondants. Le diagramme ci-dessous résume le flux de travail global utilisé pour l'identification et la quantification des protéines. Nos approches basées sur la protéomique permettent le multiplexage parallèle de la détection et de la quantification des protéines avec un degré élevé de précision et de confiance.
Diagramme global du flux de travail